Problem - Typing
there are various persistent errors in the ADL 1.4 archetypes in CKM, and presumably throughout openEHR systems. The two most common I have come across are as follows, shown as diffs, where I have made manual updates to fix the errors.
The first is creating inline ODIN (=dADL) sections for C_DV_QUANTITY when the only intention is to constraint the type to DV_QUANTITY. The correct ADL is shown in green below.

The second is the (rare) case where generic types have their internal elements set, when again, the only intent is to constrain the type (to DV_QUANTITY or DV_COUNT, in the second case). This can be replaced by the much simpler ADL shown in green.

I assume these wrong outputs are the result of never-fixed bugs in the old Archetype Editor. Hopefully these are fixed in the newer tools. Secondly, it would be good if all of the existing archetypes containing these errors could be corrected in a one-off manual upgrade. There are not many, according to the AWB:
Of course, it would be even better if we could move the archetypes and CKM to ADL2.
Problem - lack of specialisation
TB
I see groups of archetypes with commonalities, which are not modelled as specialisation hierarchies. E.g. the palpation archetypes, shown in the AWB:
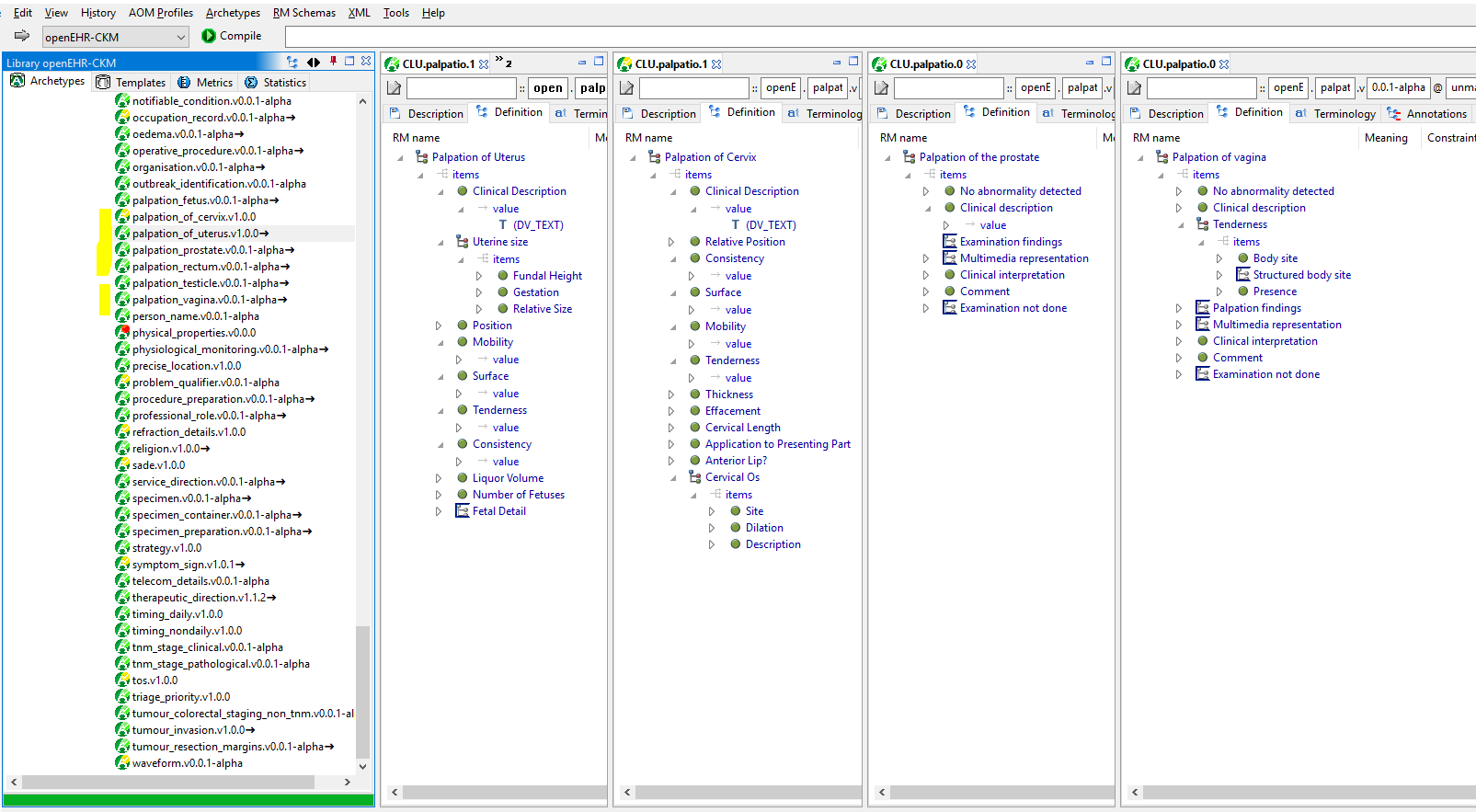
There are two timing archetypes that have a lot in common and I am sure many others, I just started looking.
The problems with not using specialisation are various:
- repeated redefinition of the same nodes in each archetype
- all of the initially duplicated elements are like a 'fork' of each other - they can all change independently, and if you change one in a certain way you (presumably) have to repeat that work N times, meaning N archetype updates, not just one
- various limitations in querying
- etc.
I guess the problematic specialisation functionality of the AE and ADL1.4 is causing this, but I see it as a very serius problem - it really limits the quality of some / ? many archetypes.
Another example:
this is exam, followed by specific exams. It seems pretty obvious that they should all be specialisations of the first. The highlighted elements show the effects of copying rather than specialising.

Similarly, exam_eye and exam_eye_anterior etc are essentialy the same except for a specialisation of the slots.
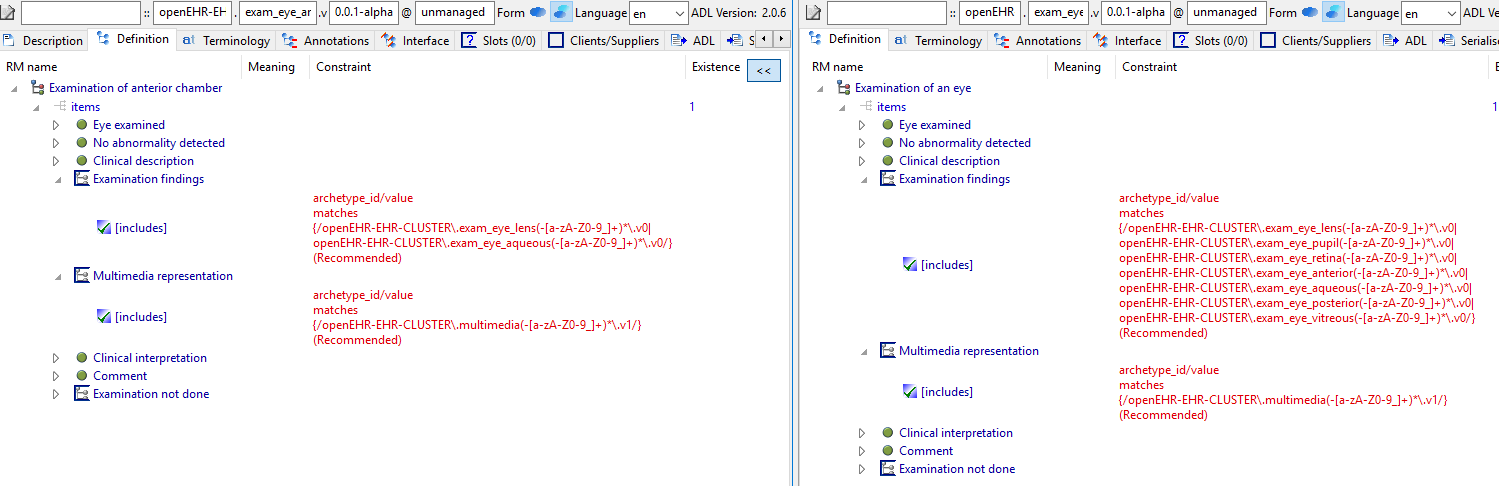
I think this style of modelling will create maintenance problems in the future (changing a common field means changing 20 archetypes potentially), translation work is multiplied, but also quite serious problems for querying right now, e.g. finding all physical exams for which 'no abnormality detected = False' (i.e. abnormal exams).
I fixed some of the exam archetypes in a area in the AWB test archetypes. It shows pretty clearly that these all should be specialisations:
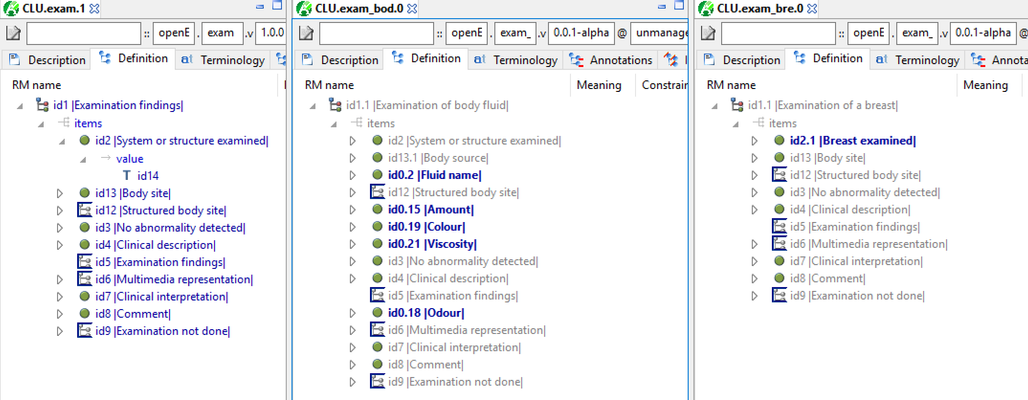
Silje
As you’re mentioning, specialisations being copies in ADL 1.4 is the main reason why we’re avoiding specialisation in general. We don’t have the tools (editors, and especially CKM) to work with ADL2.
On to your examples:
- The timing archetypes are two different concepts, timing within a day and timing outside a day. It doesn’t make sense to specialise one into the other, and I don’t think we can make abstract archetypes to make a parent “Timing” class? I’m not sure the problems with the current approach are particularly important given that we’re only talking about two archetypes.
- The exam clusters is an area where we haven’t fully agreed on a modelling approach yet. (We’re in agreement about the generic information of an examination, ie what’s in the published CLUSTER.exam archetype.)
- One approach is to make specific archetypes for each thing that’s examined. These could be copies like now, or specialisations if we had tooling for ADL2. They’d all be based on more or less the same pattern, with any additional structured elements added as requirements arise.
- Pro: Doesn’t require a terminology to make it work.
- Con: Requires a separate archetype to be created for each thing to be examined, even if there are no additional structured elements.
- Con: Hard to maintain without ADL2 specialisation (with tools).
- Another approach is to have only one examination findings CLUSTER for the generic stuff, and use terminology to specify what’s being examined. Any additional structured elements would be put in additional CLUSTERs without the generic elements, that are slotted into the examination findings CLUSTER.
- Pro: Doesn’t require us to create one archetype for each thing that can be examined, unless there are additional structured elements. This allows for quick development of new functionality with semistructured information.
- Pro: Doesn’t require specialisation to maintain.
- Con: Requires a comprehensive terminology such as SNOMED CT to work properly.
TB response:
this tools issue is many years old now. Actually, there are tools that use ADL2 - ADL-designer (internally it is AOM2), Archie (fully ADL2), and LinkEHR is fairly close. The problem is that if we stick to the old tools and methods, we are creating badly structured and modelled archetypes (in a technical sense) that are going to cause a lot of problems in the future.
The problems created by using copies are numerous:
- it's not actually clear in any case if there is a design requirement for the separation, when so many elements are the same across a related family. Future modellers will be confused with this.
- multiplies the maintenance task every time you want to do anything to a common element
- creates the risk of forking - i.e. differing changes to nodes that are meant to be the same; I already found this in the exam archetypes
- over time, or due to different people / tools etc, the codes on the nodes that are meant to be the same can differ (I already found this in the exam archetypes). This means there is no hope of making reliable AQL queries on common elements of data from that family of archetypes.
- Even if all the codes of the common elements are preserved across the separate archetypes, AQL queries for those elements are unmanageable anyway, if there are large numbers, as in the case of exam.
- creating related archetype as copies is greatly increasing the work required to convert to ADL2 in the future (hopefully very soon) - for example with these exam archetypes, I had to do an hour's work to manually convert half a dozen. But doing that properly and checking for errors, ensuring requirements are still being met etc will take significantly longer.
In the end, I think this is not good for openEHR.
The short term solution in my view is to use ADL-designer, which is internally AOM2. The exam archetypes, to take an example can be adjusted to create the correct specialisation hierarchy, even in ADL 1.4. This can be checked for correctness with the ADL workbench, and I believe ADL-designer will create the correct internal AOM2 representation. WHen changes are made in that tool, it will happen in AOM2 internally. ADL-designer can down-convert on save, to ADL1.4, so the original 'flat' form archetypes will be regenerated on output.
Silje response:
It feels to me like this discussion is about governance and modelling from my end, and modelling and implementation from your end…
If we can’t govern archetypes in ADL2 how does it matter what we model them in? Until we have a working ADL2 CKM, I don’t see how any of this would be useful. If that’s easy to do; fantastic! I’m not a developer though, so I’ll just have to wait and see.
About making an archetype abstract; I’m not sure abstract querying for timing would be at all useful. I can see its potential use for example for different types of service requests, where incidentally it also makes sense to instantiate the parent archetype. In any case, we would still need to be able to govern an abstract archetype in the CKM, and would need to be able to tag it as abstract there, to avoid people actually using it in implementations.
I think we should work to reconcile our different approaches to examination modelling before we do any significant work in changing the archetypes. If we do end up with the conclusion that at least some archetypes should be specialisations, and we have working ADL2 tools, then that rebuilding could be very useful. 😊
Wrong Terminology Mappings in Templates
BF/SLB
external terminology codes are often found encoded like;
SNOMED-CT::67101007::TX - Primærtumor kan ikke vurderes
SNOMED-CT::58790005::T0 - Primærtumor ikke påvist
SNOMED-CT::369981001::Tis - Carcinoma in situ: Intraepitelial eller infiltrerer lamina propria
SNOMED-CT::369920002::T1 - Tumor vokser inn i submucosa
SNOMED-CT::369912004::T2 - Tumor vokser inn i muscularis propria
SNOMED-CT::369921003::T3 - Tumor infiltrerer gjennom muscularis propria og inn i perikolisk vev
SNOMED-CT::369914003::T4 - Tumor vokser direkte inn i andre organer eller strukturer
What’s the standard behind this encoding format?
Is there any reference/description of this concept encoding format? (Looks like some relative of SNOMED snomed fully specified name?)
I guess it's: <terminolgyID>::<identifier>::<name> - <description>
SBL: This is the format output by Ocean Template Designer when saving a terminology value set to a text file.

TB:
Aha - in this case, the 'T0', 'TX' etc are part of the term text from SNOMED. They should not be in the Term code string. This is an error in the TD.
There is a plan to allow the format
SNOMED-CT::369920002|T1: Tumor invades submucosa|
in archetypes more or less anywhere you can currently put SNOMED-CT::369920002::T1, but not implemented yet.
SLB:
I guess this is a good idea, since it matches SNOMED’s own syntax. I suggest both ways should probably be supported for compatibility, since this syntax is used in OETs too. See attached OET from Template Designer:
<Content xsi:type="EVALUATION" archetype_id="openEHR-EHR-EVALUATION.problem_diagnosis.v1" concept_name="*Problem/Diagnosis(en)" path="/content">
<Rule path="/data[at0001]/items[at0002]">
<constraint xsi:type="textConstraint">
<includedValues>SNOMED-CT::12345::Test SNOMED term</includedValues>
</constraint>
</Rule>
</Content> |







